Molecule Tutorials - Herong's Tutorial Examples - v1.26, by Herong Yang
Export Molecule Substructure in PyMol
This section provides a tutorial on how to export a substructure identified as a selection in a molecule from PyMol.
Once a substructure is identified as a selection in PyMol, you can export the substructure to an SDF (Structure Data File) file.
1. Load the molecule from an SDF file: Molecule-HY-001.sdf:
PyMOL>delete all PyMOL>reset PyMOL>load Molecule-HY-001.sdf
2. Create the "ring" selection:
PyMOL>select ring, id 6+9+10+11+12+13 Selector: selection "ring" defined with 6 atoms.
3. Click "File > Export Molecule" from the menu. You see a dialog box.
4. Select "ring" from the "Selection" list. And click "Save" button. You see another dialog box.
5. Select "MDL SD (*.sdf *.mol)" from the format list. And enter "Benzene-Ring.sdf" as the file name.
6. Click "Save" button. The "ring" substructure is saved an SDF file called "Benzene-Ring.sdf".
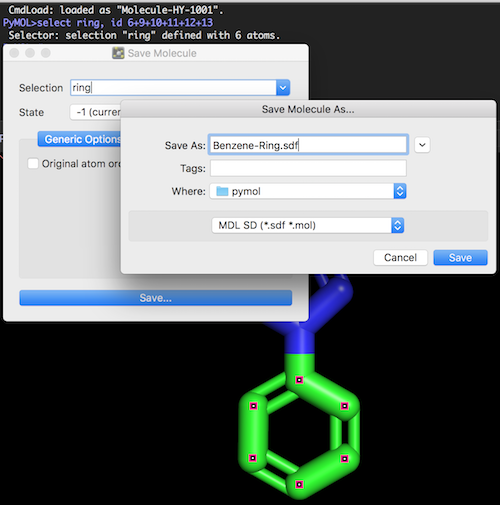
Table of Contents
Molecule Names and Identifications
Nucleobase, Nucleoside, Nucleotide, DNA and RNA
Create Selection with Mouse in PyMol
Substructure Selection Visualization in PyMol
Modify Molecule Structure in PyMol
►Export Molecule Substructure in PyMol
Create Methane Molecule in PyMol
ChEMBL Database - European Molecular Biology Laboratory
PubChem Database - National Library of Medicine
INSDC (International Nucleotide Sequence Database Collaboration)
HGNC (HUGO Gene Nomenclature Committee)