Molecule Tutorials - Herong's Tutorial Examples - v1.26, by Herong Yang
"pk1", "pk2", "pk3" and "pk4" Selections
This section provides a quick intrdocution on speical selections, pk1, pk2, pk3 and pk4, used for molecule editing functions in PyMol.
PyMol supports 4 special selections called picked atoms named as, pk1, pk2, pk3 and pk4, that are used by molecule editing functions.
The easiest way to create "pk1", "pk2", "pk3" and "pk4" selections is to use the mouse click in "Editing" mode.
1. Load the molecule from Molecule-HY-001.sdf.
PyMOL>delete all PyMOL>reset PyMOL>load Molecule-HY-001.sdf
2. Click "Mouse Mode" to set it to "3-Button Editing" in the mouse control area near the bottom right corner of the viewer window.
3. Click an atom on the molecule. You see pk1 selection created for the selected atom.
4. Click 3 more atoms. You see all 4 special selections, pk1, pk2, pk3 and pk4, created and identified with different markers.
You can remove any pk* selection by click it again.
See next tutorials on how to use those 4 picked atoms (pk* selections).
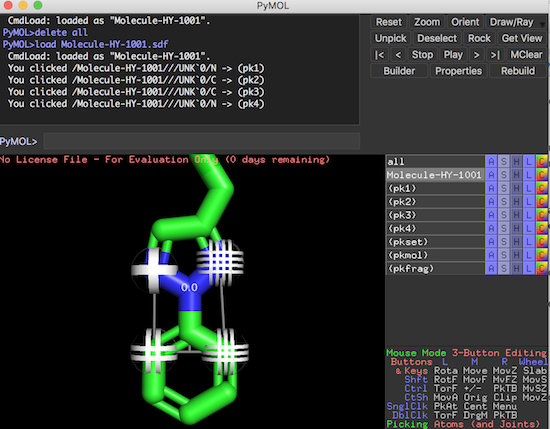
Table of Contents
Molecule Names and Identifications
Nucleobase, Nucleoside, Nucleotide, DNA and RNA
►"pk1", "pk2", "pk3" and "pk4" Selections
"edit id n1, id n2, id n3, id n4" Commands
"remove pk*" and "remove_picked" Commands
"unbond pk1, pk2" and "bond pk1, pk2" Commands
"replace new_atom, ..." Replace pk1 with New Atom
"attach new_atom, ..." Attach to pk1 with New Atom
Build Alcohol Molecule with PyMol
ChEMBL Database - European Molecular Biology Laboratory
PubChem Database - National Library of Medicine
INSDC (International Nucleotide Sequence Database Collaboration)
HGNC (HUGO Gene Nomenclature Committee)