Molecule Tutorials - Herong's Tutorial Examples - v1.26, by Herong Yang
Build Alcohol Molecule with PyMol
This section provides a tutorial on how to build an alcohol molecule with PyMol commands.
PyMol also have a command add hydrogen atoms to the molecule structure if there are not include with the "h_add" command.
Together with "h_add" and "replace" commands, we have build an alcohol molecule from a single carbon atom.
delete all reset # load the carbon atom from an SDF file load Carbon-Atom.sdf hide all # add hydrogen atoms to form the methane molecule h_add show sticks # replace one hydrogen atom to carbon edit id 2 replace C, 1, 1 # replace another hydrogon atom to oxygen edit id 3 replace O, 1, 1 # save the molecule to an SDF file save Alcohol-Molecule.sdf, all, -1, sdf
The result is an alcohol molecule. Some hydrogen atoms are not located in correct angles. I can fix them later.
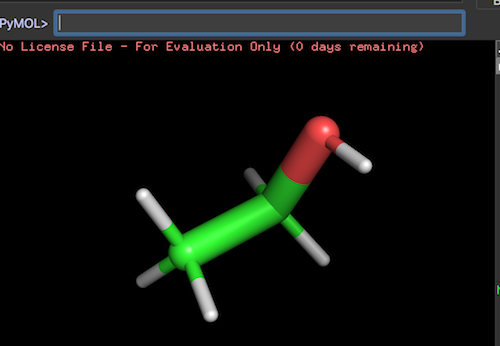
Table of Contents
Molecule Names and Identifications
Nucleobase, Nucleoside, Nucleotide, DNA and RNA
"pk1", "pk2", "pk3" and "pk4" Selections
"edit id n1, id n2, id n3, id n4" Commands
"remove pk*" and "remove_picked" Commands
"unbond pk1, pk2" and "bond pk1, pk2" Commands
"replace new_atom, ..." Replace pk1 with New Atom
"attach new_atom, ..." Attach to pk1 with New Atom
►Build Alcohol Molecule with PyMol
ChEMBL Database - European Molecular Biology Laboratory
PubChem Database - National Library of Medicine
INSDC (International Nucleotide Sequence Database Collaboration)
HGNC (HUGO Gene Nomenclature Committee)