Molecule Tutorials - Herong's Tutorial Examples - v1.26, by Herong Yang
"unbond pk1, pk2" and "bond pk1, pk2" Commands
This section provides a tutorial on how to unbond or bond atoms from a molecule in PyMol using picked atoms as special selections, pk1, pk2, pk3 and pk4.
You can unbond or bond atoms picked atoms as pk1, pk2, pk3 and pk4 selections from the molecule structure. For example:
PyMOL># remove the bond between first two picked atoms: PyMOL>unbond pk1, pk2 PyMOL># add a bond between first two picked atoms PyMOL>bond pk1, pk2
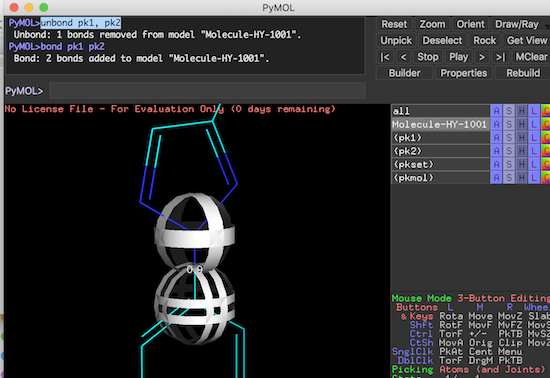
Table of Contents
Molecule Names and Identifications
Nucleobase, Nucleoside, Nucleotide, DNA and RNA
"pk1", "pk2", "pk3" and "pk4" Selections
"edit id n1, id n2, id n3, id n4" Commands
"remove pk*" and "remove_picked" Commands
►"unbond pk1, pk2" and "bond pk1, pk2" Commands
"replace new_atom, ..." Replace pk1 with New Atom
"attach new_atom, ..." Attach to pk1 with New Atom
Build Alcohol Molecule with PyMol
ChEMBL Database - European Molecular Biology Laboratory
PubChem Database - National Library of Medicine
INSDC (International Nucleotide Sequence Database Collaboration)
HGNC (HUGO Gene Nomenclature Committee)