Molecule Tutorials - Herong's Tutorial Examples - v1.26, by Herong Yang
"show lines" Presentation Command
This section provides a tutorial on how the 'show' command to turn on the 'lines' representation of the loaded molecule structure in PyMol.
Once you have turned and moved your camera to a good angle and a good distance in PyMol, you can use "hide" and "show" commands to change the representation style of the molecule structure.
The simplest representation is called "lines". You can turn it on and off with these commands:
show lines hide lines
Let's load a molecule and show it in "lines" representation:
PyMOL># open a log file to record commands PyMOL>log_open Representation-Show-Lines.pml PyMOL># delete all objects PyMOL>delete all PyMOL># reset viewing parameters to default values PyMOL>reset PyMOL># load my molecule from a file PyMOL>load Molecule-HY-001.sdf CmdLoad: loaded as "Molecule-HY-001". PyMOL># hide all representations PyMOL>hide all PyMOL># show "lines" representation PyMOL>show lines PyMOL># move the camera backward to scale down the representation PyMOL>move z, -5 PyMOL># close the log file PyMOL>log_close PyMOL># re-run all commands in the log file PyMOL>@Representation-Show-Lines.pml
The picture below shows the "lines" representation of a molecule structure:
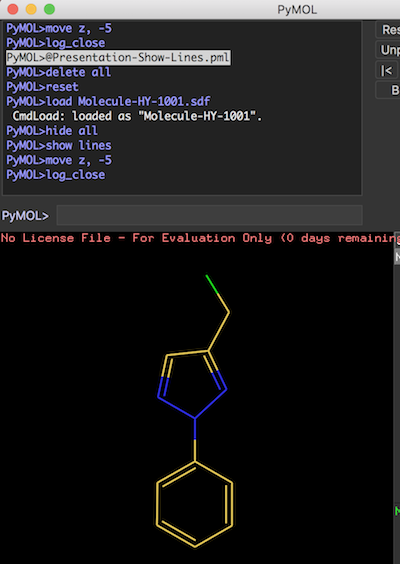
Table of Contents
Molecule Names and Identifications
Nucleobase, Nucleoside, Nucleotide, DNA and RNA
Load Molecule from File into PyMol
Virtual Trackball Rotation on PyMol
"load" and "delete" Commands on PyMol
"log_open" and "log_close" Commands on PyMol
Model Space and Camera Space on PyMol
"get_view" and "set_view" on PyMol
View Parameters Auto Adjusted on PyMol
Rotation with Transformation Matrix
Difference of "turn" and "rotate" Commands
Difference of "move" and "translate" Commands
"center", "zoom" and "reset" Commands
Model-to-Camera Space Coordinates Mapping
Camera-to-Model Space Coordinates Mapping
►"show lines" Presentation Command
"show sticks" Presentation Command
"show spheres" Presentation Command
"show surface" Presentation Command
"show mesh" Presentation Command
ChEMBL Database - European Molecular Biology Laboratory
PubChem Database - National Library of Medicine
INSDC (International Nucleotide Sequence Database Collaboration)
HGNC (HUGO Gene Nomenclature Committee)