Molecule Tutorials - Herong's Tutorial Examples - v1.26, by Herong Yang
Zoom In and Out on PyMol
This section provides a tutorial on how to zoom in and out the viewing window on PyMol.
Once the molecule structure is rotated to the ideal direction, you can zoom in the viewing window to see more details if the molecule structure is too large using the right mouse button or the two-finger spread gesture.
You can also zoom out the viewing window by reverse mouse or figure controls.
For example, if you want to zoom in the viewing window, place two figures on the touchpad, and spread them apart. You see the molecule structure image is enlarged, as if the viewing window is zoomed in at the center of the screen.
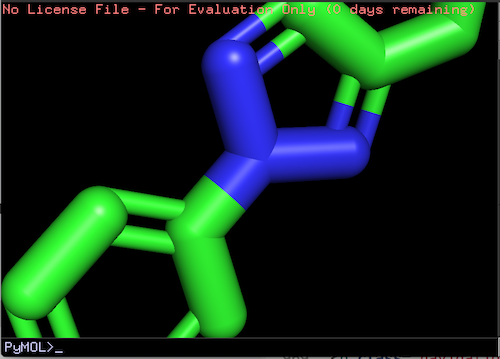
Table of Contents
Molecule Names and Identifications
Nucleobase, Nucleoside, Nucleotide, DNA and RNA
Load Molecule from File into PyMol
Virtual Trackball Rotation on PyMol
"load" and "delete" Commands on PyMol
"log_open" and "log_close" Commands on PyMol
Model Space and Camera Space on PyMol
"get_view" and "set_view" on PyMol
View Parameters Auto Adjusted on PyMol
Rotation with Transformation Matrix
Difference of "turn" and "rotate" Commands
Difference of "move" and "translate" Commands
"center", "zoom" and "reset" Commands
Model-to-Camera Space Coordinates Mapping
Camera-to-Model Space Coordinates Mapping
"show lines" Presentation Command
"show sticks" Presentation Command
"show spheres" Presentation Command
"show surface" Presentation Command
"show mesh" Presentation Command
ChEMBL Database - European Molecular Biology Laboratory
PubChem Database - National Library of Medicine
INSDC (International Nucleotide Sequence Database Collaboration)
HGNC (HUGO Gene Nomenclature Committee)